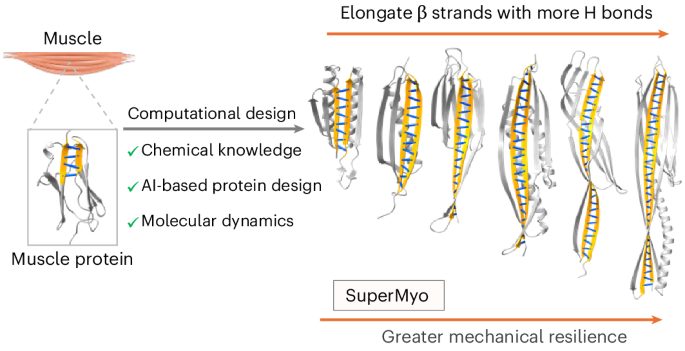
Computational design of superstable proteins through maximized hydrogen bonding
Rief, M., Gautel, M., Oesterhelt, F., Fernandez, J. M. & Gaub, H. E. Reversible unfolding of individual titin immunoglobulin domains by AFM. Science 276, 1109–1112 (1997).
Google Scholar
Li, H. B. et al. Reverse engineering of the giant muscle protein titin. Nature 418, 998–1002 (2002).
Google Scholar
Keten, S., Xu, Z., Ihle, B. & Buehler, M. J. Nanoconfinement controls stiffness, strength and mechanical toughness of β-sheet crystals in silk. Nat. Mater. 9, 359–367 (2010).
Google Scholar
Nelson, R. et al. Structure of the cross-β spine of amyloid-like fibrils. Nature 435, 773–778 (2005).
Google Scholar
Johnson, C. P., Tang, H.-Y., Carag, C., Speicher, D. W. & Discher, D. E. Forced unfolding of proteins within cells. Science 317, 663–666 (2007).
Google Scholar
Bustamante, C., Chemla, Y. R., Forde, N. R. & Izhaky, D. Mechanical processes in biochemistry. Annu. Rev. Biochem. 73, 705–748 (2004).
Google Scholar
Roca-Cusachs, P., Conte, V. & Trepat, X. Quantifying forces in cell biology. Nat. Cell Biol. 19, 742–751 (2017).
Google Scholar
Hoffmann, T., Tych, K. M., Hughes, M. L., Brockwell, D. J. & Dougan, L. Towards design principles for determining the mechanical stability of proteins. Phys. Chem. Chem. Phys. 15, 15767–15780 (2013).
Google Scholar
Milles, L. F., Schulten, K., Gaub, H. E. & Bernardi, R. C. Molecular mechanism of extreme mechanostability in a pathogen adhesin. Science 359, 1527–1533 (2018).
Google Scholar
Craig, L., Forest, K. T. & Maier, B. Type IV pili: dynamics, biophysics and functional consequences. Nat. Rev. Microbiol. 17, 429–440 (2019).
Google Scholar
Tskhovrebova, L. & Trinick, J. Titin: properties and family relationships. Nat. Rev. Mol. Cell. Bio. 4, 679–689 (2003).
Google Scholar
Labeit, S. & Kolmerer, B. Titins: giant proteins in charge of muscle ultrastructure and elasticity. Science 270, 293–296 (1995).
Google Scholar
Brockwell, D. J. et al. Pulling geometry defines the mechanical resistance of a beta-sheet protein. Nat. Struct. Mol. Biol. 10, 731–737 (2003).
Google Scholar
Dietz, H. & Rief, M. Exploring the energy landscape of GFP by single-molecule mechanical experiments. Proc. Natl Acad. Sci. USA 101, 16192–16197 (2004).
Google Scholar
Rico, F., Gonzalez, L., Casuso, I., Puig-Vidal, M. & Scheuring, S. High-speed force spectroscopy unfolds titin at the velocity of molecular dynamics simulations. Science 342, 741–743 (2013).
Google Scholar
Bull, M. S., Sullan, R. M. A., Li, H. & Perkins, T. T. Improved single molecule force spectroscopy using micromachined cantilevers. ACS Nano 8, 4984–4995 (2014).
Google Scholar
Brockwell, D. J. et al. Mechanically unfolding the small, topologically simple protein L. Biophys. J. 89, 506–519 (2005).
Google Scholar
Bertz, M. & Rief, M. Ligand binding mechanics of maltose binding protein. J. Mol. Biol. 393, 1097–1105 (2009).
Google Scholar
Cao, Y., Yoo, T. & Li, H. Single molecule force spectroscopy reveals engineered metal chelation is a general approach to enhance mechanical stability of proteins. Proc. Natl Acad. Sci. USA 105, 11152–11157 (2008).
Google Scholar
López-García, P. et al. Fortified coiled coils: enhancing mechanical stability with lactam or metal staples. Angew. Chem. Int. Ed. 60, 232–236 (2021).
Google Scholar
Müller, D. J. et al. Atomic force microscopy-based force spectroscopy and multiparametric imaging of biomolecular and cellular systems. Chem. Rev. 121, 11701–11725 (2021).
Google Scholar
Li, H., Carrion-Vazquez, M., Oberhauser, A. F., Marszalek, P. E. & Fernandez, J. M. Point mutations alter the mechanical stability of immunoglobulin modules. Nat. Struct. Mol. Biol. 7, 1117–1120 (2000).
Google Scholar
Ferruz, N. & Höcker, B. Dreaming ideal protein structures. Nat. Biotechnol. 40, 171–172 (2022).
Google Scholar
Huang, B. et al. A backbone-centred energy function of neural networks for protein design. Nature 602, 523–528 (2022).
Google Scholar
Kortemme, T. De novo protein design—from new structures to programmable functions. Cell 187, 526–544 (2024).
Google Scholar
Dauparas, J. et al. Atomic context-conditioned protein sequence design using LigandMPNN. Nat. Methods 22, 717–723 (2025).
Google Scholar
Schissel, C. K. et al. Deep learning to design nuclear-targeting abiotic miniproteins. Nat. Chem. 13, 992–1000 (2021).
Google Scholar
Chen, K.-Y. et al. Computational design of highly signalling-active membrane receptors through solvent-mediated allosteric networks. Nat. Chem. 17, 429–438 (2025).
Google Scholar
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Google Scholar
Lin, Z. et al. Evolutionary-scale prediction of atomic-level protein structure with a language model. Science 379, 1123–1130 (2023).
Google Scholar
Watson, J. L. et al. De novo design of protein structure and function with RFdiffusion. Nature 620, 1089–1100 (2023).
Google Scholar
Dauparas, J. et al. Robust deep learning–based protein sequence design using ProteinMPNN. Science 378, 49–56 (2022).
Google Scholar
Sumida, K. H. et al. Improving protein expression, stability, and function with ProteinMPNN. J. Am. Chem. Soc. 146, 2054–2061 (2024).
Google Scholar
Ni, B., Kaplan, D. L. & Buehler, M. J. ForceGen: end-to-end de novo protein generation based on nonlinear mechanical unfolding responses using a language diffusion model. Sci. Adv. 10, eadl4000 (2024).
Google Scholar
Sikora, M. & Cieplak, M. Mechanical stability of multidomain proteins and novel mechanical clamps. Proteins 79, 1786–1799 (2011).
Google Scholar
Karplus, M. & McCammon, J. A. Molecular dynamics simulations of biomolecules. Nat. Struct. Mol. Biol. 9, 646–652 (2002).
Google Scholar
Van Der Spoel, D. et al. GROMACS: fast, flexible, and free. J. Comput. Chem. 26, 1701–1718 (2005).
Google Scholar
Bernardi, R. C. et al. Mechanisms of nanonewton mechanostability in a protein complex revealed by molecular dynamics simulations and single-molecule force spectroscopy. J. Am. Chem. Soc. 141, 14752–14763 (2019).
Google Scholar
Day, R., Bennion, B. J., Ham, S. & Daggett, V. Increasing temperature accelerates protein unfolding without changing the pathway of unfolding. J. Mol. Biol. 322, 189–203 (2002).
Google Scholar
Mayor, U., Johnson, C. M., Daggett, V. & Fersht, A. R. Protein folding and unfolding in microseconds to nanoseconds by experiment and simulation. Proc. Natl Acad. Sci. USA 97, 13518–13522 (2000).
Google Scholar
Huang, J. et al. CHARMM36m: an improved force field for folded and intrinsically disordered proteins. Nat. Methods 14, 71–73 (2017).
Google Scholar
Lu, H., Isralewitz, B., Krammer, A., Vogel, V. & Schulten, K. Unfolding of titin immunoglobulin domains by steered molecular dynamics simulation. Biophys. J. 75, 662–671 (1998).
Google Scholar
Chen, H. et al. Dynamics of equilibrium folding and unfolding transitions of titin immunoglobulin domain under constant forces. J. Am. Chem. Soc. 137, 3540–3546 (2015).
Google Scholar
Lu, H. & Schulten, K. The key event in force-induced unfolding of titin’s immunoglobulin domains. Biophys. J. 79, 51–65 (2000).
Google Scholar
Klimov, D. K. & Thirumalai, D. Native topology determines force-induced unfolding pathways in globular proteins. Proc. Natl Acad. Sci. USA 97, 7254–7259 (2000).
Google Scholar
Yu, M., Lu, J.-H., Le, S. & Yan, J. Unexpected low mechanical stability of titin I27 domain at physiologically relevant temperature. J. Phys. Chem. Lett. 12, 7914–7920 (2021).
Google Scholar
Mirdita, M. et al. ColabFold: making protein folding accessible to all. Nat. Methods 19, 679–682 (2022).
Google Scholar
Phillips, J. C. et al. Scalable molecular dynamics on CPU and GPU architectures with NAMD. J. Chem. Phys. 153, 044130 (2020).
Google Scholar
Gräter, F., Shen, J., Jiang, H., Gautel, M. & Grubmüller, H. Mechanically induced titin kinase activation studied by force-probe molecular dynamics simulations. Biophys. J. 88, 790–804 (2005).
Google Scholar
Lee, J. et al. CHARMM-GUI input generator for NAMD, GROMACS, AMBER, OpenMM, and CHARMM/OpenMM simulations using the CHARMM36 additive force field. J. Chem. Theory Comput. 12, 405–413 (2015).
Google Scholar
Kim, M., Kim, E., Lee, S., Kim, J. S. & Lee, S. New method for constant-NPT molecular dynamics. J. Phys. Chem. A 123, 1689–1699 (2019).
Google Scholar
Zhang, P., Wang, D., Yang, W. T. & Marszalek, P. E. Piecewise all-atom SMD simulations reveal key secondary structures in luciferase unfolding pathway. Biophys. J. 119, 2251–2261 (2020).
Google Scholar
Khare, E. et al. Discovering design principles of collagen molecular stability using a genetic algorithm, deep learning, and experimental validation. Proc. Natl Acad. Sci. USA 119, e2209524119 (2022).
Google Scholar
Müller, D. J. & Dufrene, Y. F. Atomic force microscopy as a multifunctional molecular toolbox in nanobiotechnology. Nat. Nanotechnol. 3, 261–269 (2008).
Google Scholar
Deng, Y. et al. Enzymatic biosynthesis and immobilization of polyprotein verified at the single-molecule level. Nat. Commun. 10, 2775–2785 (2019).
Google Scholar
Shi, S. et al. Combination of click chemistry and enzymatic ligation for stable and efficient protein immobilization for single-molecule force spectroscopy. CCS Chem 4, 598–604 (2022).
Google Scholar
Ott, W. et al. Elastin-like polypeptide linkers for single-molecule force spectroscopy. ACS Nano 11, 6346–6354 (2017).
Google Scholar
Marko, J. F. & Siggia, E. D. Stretching DNA. Macromolecules 28, 8759–8770 (1995).
Google Scholar
Cao, Y., Lam, C., Wang, M. & Li, H. Nonmechanical protein can have significant mechanical stability. Angew. Chem. Int. Ed. 45, 642–645 (2006).
Google Scholar
Carrion-Vazquez, M. et al. Mechanical and chemical unfolding of a single protein: a comparison. Proc. Natl Acad. Sci. USA 96, 3694–3699 (1999).
Google Scholar
Greenfield, N. J. Using circular dichroism spectra to estimate protein secondary structure. Nat. Protoc. 1, 2876–2890 (2007).
Google Scholar
Baran, M. C., Huang, Y. J., Moseley, H. N. & Montelione, G. T. Automated analysis of protein NMR assignments and structures. Chem. Rev. 104, 3541–3556 (2004).
Google Scholar
Merkel, R., Nassoy, P., Leung, A., Ritchie, K. & Evans, E. Energy landscapes of receptor–ligand bonds explored with dynamic force spectroscopy. Nature 397, 50–53 (1999).
Google Scholar
de Gennes, P.-G. Maximum pull out force on DNA hybrids. C. R. Acad. Sci. IV 2, 1505–1508 (2001).
Politou, A. S., Thomas, D. J. & Pastore, A. The folding and stability of titin immunoglobulin-like modules, with implications for the mechanism of elasticity. Biophys. J. 69, 2601–2610 (1995).
Google Scholar
Joosten, R. P. et al. A series of PDB related databases for everyday needs. Nucleic Acids Res. 39, D411–D419 (2010).
Google Scholar
Mout, R. et al. De novo design of modular protein hydrogels with programmable intra-and extracellular viscoelasticity. Proc. Natl Acad. Sci. USA 121, e2309457121 (2024).
Google Scholar
Liao, H. et al. Data-driven de novo design of super-adhesive hydrogels. Nature 644, 89–95 (2025).
Google Scholar
Lv, S. et al. Designed biomaterials to mimic the mechanical properties of muscles. Nature 465, 69–73 (2010).
Google Scholar
Doolan, J. A. et al. Next-generation protein-based materials capture and preserve projectiles from supersonic impacts. Nat. Nanotechnol. 18, 1060–1066 (2023).
Google Scholar
Sun, F., Zhang, W.-B., Mahdavi, A., Arnold, F. H. & Tirrell, D. A. Synthesis of bioactive protein hydrogels by genetically encoded SpyTag-SpyCatcher chemistry. Proc. Natl Acad. Sci. USA 111, 11269–11274 (2014).
Google Scholar
Ko, J. H. & Maynard, H. D. A guide to maximizing the therapeutic potential of protein–polymer conjugates by rational design. Chem. Soc. Rev. 47, 8998–9014 (2018).
Google Scholar
Gao, X., Fang, J., Xue, B., Fu, L. & Li, H. Engineering protein hydrogels using SpyCatcher-SpyTag chemistry. Biomacromolecules 17, 2812–2819 (2016).
Google Scholar
Yang, R. et al. Engineering a catalytically efficient recombinant protein ligase. J. Am. Chem. Soc. 139, 5351–5358 (2017).
Google Scholar
Hutter, J. L. & Bechhoefer, J. Calibration of atomic-force microscope tips. Revi. Sci. Instrum. 64, 1868–1873 (1993).
Google Scholar
Delaglio, F. et al. NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J. Biomol. NMR 6, 277–293 (1995).
Google Scholar
Lee, W., Tonelli, M. & Markley, J. L. NMRFAM-SPARKY: enhanced software for biomolecular NMR spectroscopy. Bioinformatics 31, 1325–1327 (2015).
Google Scholar
Lee, W. et al. I-PINE web server: an integrative probabilistic NMR assignment system for proteins. J. Biomol. NMR 73, 213–222 (2019).
Google Scholar
Zhang, Y. et al. Advances in transdermal insulin delivery. Adv. Drug Deliv. Rev. 139, 51–70 (2019).
Google Scholar
Güntert, P. Automated NMR structure calculation with CYANA. Protein NMR Tech. 278, 353–378 (2004).
Google Scholar
Bhattacharya, A., Tejero, R. & Montelione, G. T. Evaluating protein structures determined by structural genomics consortia. Proteins 66, 778–795 (2007).
Google Scholar
link






