Structure-based design of potent and selective inhibitors targeting RIPK3 for eliminating on-target toxicity in vitro
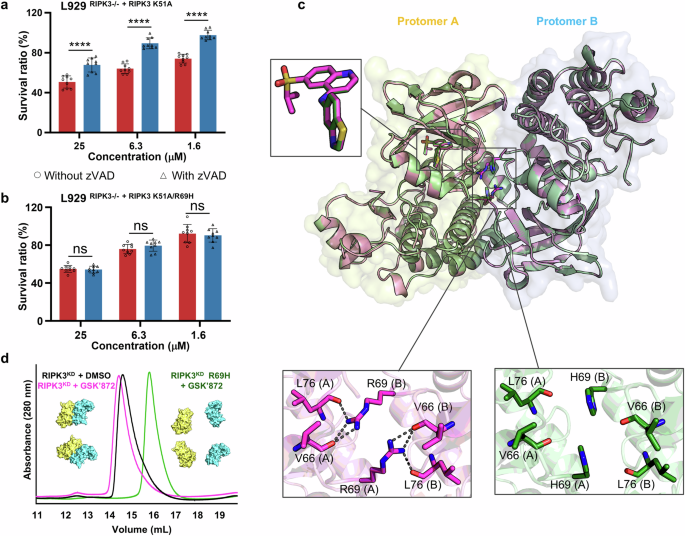
Co-crystal structures of mRIPK3KD and mRIPK3KD R69H in complex with GSK’872
A previous study by Raju et al. suggestes that the R69H mutation eliminates GSK’872-induced apoptosis by dissociating RIPK3KD dimers into monomers34. In this context, we first examine whether the R69H mutation could prevent on-target apoptosis in L929 cells, a widely used cell line for studying RIPK3 inhibitors. Our results show that GSK’872, at low micromolar concentrations (e.g., 1.6 μM), significantly reduces the viability of cells expressing the ATPase-inactive RIPK3 K51A mutant (Fig. 1a). Importantly, the loss of cell viability is reversed by the addition of zVAD, a pan-caspase inhibitor that prevents apoptosis, suggesting that the observed cell death is a result of apoptosis induced by GSK’872. However, GSK’872 does not cause such apoptosis in cells expressing RIPK3 with the double mutation of K51A and R69H (Fig. 1b). This confirms that the R69H mutation can rescue cells from apoptosis triggered by GSK’872.

a, b Viability of L929 cells with RIPK3 knockout (RIPK3-/-) and overexpressed RIPK3 K51A (a) or RIPK3 K51A/R69H (b), treated with GSK’872 with (open triangle) and without (open circle) zVAD. c Superimposition of two co-crystal structures of mRIPK3KD (magenta cartoons) and mRIPK3KD R69H (green cartoons) in complex with GSK’872 with enlarged views of bound GSK’872 and R69 or H69 with its surroundings. Hydrogen bonds are represented by black dashed lines. d Overlaid size-exclusion chromatograms (Superdex 200) of mRIPK3KD with DMSO (black lines), mRIPK3KD with GSK’872 (magenta lines), and mRIPK3KD R69H with GSK’872 (green lines). The protein is concentrated to 10 mg/mL and GSK’872 is added to a final compound concentration of 1 mM. Three independent experiments are performed for (a, b), each with three technical replicates, and data are shown as mean ± SD. The data shown in d represent one of three independent experiments, all of which yield similar results. Statistical differences are determined by multiple t-tests. **q < 0.01, ***q < 0.001, q < 0.0001. Source data are provided as a Source Data file.
Subsequently, to investigate whether the R69H mutation affects the binding mode of the inhibitor, we determine co-crystal structures of mouse RIPK3 kinase domain (mRIPK3KD) and mRIPK3KD R69H in complex with GSK’872. Superimposition of the two crystal structures shows that the bound conformations of GSK’872 and overall structures (backbone) of the protein are almost identical in both complexes (Fig. 1c). This suggests that the protective effect of the R69H mutation against inhibitor-induced on-target toxicity is not resulting from altering the inhibitor’s binding mode or the overall structure of the kinase domain.
We further examine interactions of R69 with its surroundings in the complex structure of mRIPK3KD bound with GSK’872. It is found that R69 is located at the dimer interface of the kinase domain, where it forms hydrogen bonds with the main chain of V66 and L76 from another protomer (Fig. 1c). Mutation of R69 to H69 disrupts these hydrogen-bonding interactions (Fig. 1c), which provides a structural basis for understanding how the R69H mutation impairs dimerization of mRIPK3KD. In addition, we conduct size-exclusion chromatography experiments. The results show that both apo mRIPK3KD and its complex with GSK’872 elute at a position corresponding to the molecular weight of a dimer (Fig. 1d). In contrast, the peak position for eluted mRIPK3KD R69H in complex with GSK’872 aligns with a monomer (Fig. 1d). Taken together, these results demonstrate that the R69H mutation promotes the disassembly of the mRIPK3KD dimers into monomers by disrupting the dimer interface of the kinase domain.
Structure-based design of inhibitors to mimic the R69H mutation
The co-crystal structure of mRIPK3KD in complex with GSK’872 shows that R69 is located behind an αC-helix, while the thiazole group of GSK’872 is well positioned on the other side of the αC-helix (Fig. 2a). In order to mimic the R69H mutation, we introduce hydrophobic groups on the thiazole group of GSK’872 to interfere with the αC-helix by forming direct interactions with multiple hydrophobic amino acids of the helix (Fig. 2a). As a result, compounds LK01001, LK01002, and LK01003 are designed, synthesized, and evaluated by the ADP-Glo assay, demonstrating significant inhibition against the kinase activity of RIPK3 with IC50 values of 11, 3, and 2 nM, respectively (Fig. 2b, c). The potency of these three compounds is comparable or superior to that of GSK’872, which is also tested as a reference compound and has an IC50 value of 6 nM.

a Strategy for the design of R69H-mimicking inhibitors based on GSK’872. b Chemical structures of inhibitors with their IC50 values determined by the ADP-Glo assay and EC50 values against TNF-α-induced necroptosis in L929 and HT29 cell lines determined by the CCK-8 assay. c–e Representative inhibitory profiles of GSK’872 (magenta solid circle), LK01001 (green solid triangle), LK01002 (blue solid inverted triangle), LK01003 (purple solid diamond), and LK01045 (cyan open inverted triangle) against mRIPK3KD in the ADP-Glo assay (c), and against TNF-α-induced necroptosis in L929 (d) and HT29 (e) cells. In L929 cells, necroptosis is induced by treatment with mouse TNF-α (20 ng/mL) and zVAD (10 μM). Necroptosis in HT29 cells is induced by treatment with zVAD (20 μM), human TNF-α (40 ng/mL), and Smac mimetic SM-164 (100 nM). f, g Western blot analysis of RIPK3 and MLKL phosphorylation in L929 (f) and HT29 (g) cells with or without RIPK3 inhibitors. Cells are treated with inhibitors at concentrations of 3 and 10 µM, followed by induction of necroptosis for 3 hours. h Overlaid size-exclusion chromatograms (Superdex 200) of mRIPK3KD with GSK’872 (magenta lines), LK01002 (blue lines), LK01003 (purple lines), and LK01045 (cyan lines), alongside mRIPK3KD R69H mutant with GSK’872 (green lines). The protein is concentrated to 10 mg/mL and the inhibitors are added to a final compound concentration of 1 mM. The data in panel c are plotted as the mean ± SD of triplicate wells. Three independent experiments are performed for (d, e), each with three technical replicates, and data are shown as mean ± SD. Data shown in panels (f–h) represent one of the three independent experiments, all of which showed similar results. Source data are provided as a Source Data file.
Necroptosis inhibition tests on these compounds are subsequently conducted on L929 and HT29 cell lines, which are widely used for the assessment of cellular necroptosis and potency of RIPK3 inhibitors. Necroptosis in L929 cells is induced using mouse TNF-α and zVAD. The anti-necroptotic activities of the compounds in L929 cells are determined with an EC50L929 value of 7.0 μM for LK01001, 4.8 μM for LK01002, and 3.7 μM for LK01003 (Fig. 2b, d), similar to the EC50L929 value of GSK’872 (3.6 μM). In HT29 cells, necroptosis is induced using zVAD, human TNF-α, and Smac mimetic SM-164. The EC50HT29 value of LK01001, LK01002, LK01003, and GSK’872 is 4.1, 2.0, 0.85, and 7.9 μM, respectively (Fig. 2b, e). This indicates that LK01002 and LK01003 are much more potent than GSK’872 to block necroptosis in HT29 cells. Given the higher potency of LK01002 over its enantiomer LK01001, shown in both the ADP-Glo assay and necroptosis inhibition tests in HT29 cells, we focused on LK01002 and LK01003 for the subsequent experiments. Western blot analysis confirms that LK01002 and LK01003 effectively inhibit RIPK3 and MLKL phosphorylation induced by TNF-α in both L929 and HT29 cells (Fig. 2f, g).
In addition, we introduce an oxyethanol group onto the quinoline ring of LK01003 that is facing the solvent to improve the solubility of the compound. This modification results in the development of LK01045 (Fig. 2b), which also demonstrates strong inhibition against RIPK3 kinase activity with an IC50 value of 16 nM (Fig. 2b, c). More importantly, LK01045 shows improved inhibition against TNF-α-induced necroptosis in both L929 and HT29 cells, with EC50 values of 0.61 and 0.16 µM, respectively (Fig. 2b, d, e). These EC50 values represent approximately 6-fold and 5-fold stronger potency compared to LK01003, and approximately 6-fold and 49-fold stronger potency compared to GSK’872. Western blot analysis further confirms that LK01045 effectively inhibits RIPK3 and MLKL phosphorylation induced by TNF-α in both L929 and HT29 cells (Fig. 2f, g). Together, these results show the potent activity of LK01002, LK01003, and LK01045 in preventing cell necroptosis by inhibiting the kinase activity of RIPK3.
Further investigation is carried out to evaluate the potential of LK01002, LK01003, and LK01045 to mimic the effects of the R69H mutation by examining the impact of these three compounds on the dimerization of the RIPK3 kinase domain. As expected, the binding of LK01002, LK01003, or LK01045 disrupts the dimerization of mRIPK3KD, as evidenced by the delayed peak positions in the size-exclusion chromatography, which overlap well with that of the mRIPK3KD R69H bound with GSK’872 (Fig. 2h). These results suggest that these three compounds effectively reproduce the R69H mutation’s ability to impede the dimer formation of the kinase domain.
Moreover, in the cell viability assay, toxicity due to on-target apoptosis is observed with GSK’872 at a concentration of 3.1 μM (Fig. 3a). However, none of LK01002, LK01003, and LK01045 exhibits on-target toxicity at concentrations up to 50 μM (Fig. 3b-d). Such a significant increase in the tolerated concentration level represents a breakthrough in our attempts to prevent on-target toxicity induced by the inhibitors. We further tested these compounds in L929 cells with RIPK3 knockout (L929RIPK3−/−). The CC50 value of GSK’872 significantly increases from 6.3 μM in L929 to 22 μM in L929RIPK3−/− cells (Fig. 3a, e), indicating reduced toxicity in the absence of RIPK3. In contrast, the CC50 values of LK01002, LK01003, and LK01045 in both L929 and L929RIPK3−/− cells remain largely unchanged (Fig. 3b-d, f-h), further demonstrating that these compounds do not cause on-target toxicity. We also tested LK01002, LK01003, and LK01045 in L929 cells expressing the K51A/R69H mutants and found that their CC50 values are almost identical to those in the K51A mutant cell line, further supporting the conclusion that they do not induce on-target toxicity (Supplementary Fig. 1). In addition, GSK’872 activates caspase 3 at a concentration of 5 μM (Fig. 3i). In stark contrast, none of LK01002, LK01003, and LK01045 triggers caspase 3 activation even at 50 μM (Fig. 3i), which is approximately 10, 14, and 82 times their respective EC50 values in L929 cells. Moreover, immunoprecipitation analysis reveals that GSK’872 enhances the interaction between RIPK3 and RIPK1 (Fig. 3j), thereby activating the apoptotic pathway and leading to the cleavage of caspase 8 and caspase 3 (Fig. 3k). In contrast, these three compounds neither increase the interaction of RIPK3 with RIPK1 (Fig. 3j) nor activate the apoptotic pathway (Fig. 3k). Accordingly, the absence of increased RIPK3-RIPK1 interaction and the lack of activation of apoptosis-related proteins when treated with LK01002, LK01003, and LK01045 lent strong support to the observation that the three compounds can potently inhibit RIPK3 without triggering the undesired apoptotic signaling. Also, the successful discovery of this new series of inhibitors suggests that mimicking the effect of R69H mutation offers a promising strategy for the identification of new RIPK3 inhibitors without on-target toxicity. Although LK01045 shows a more potent EC50 value compared to LK01003, subsequent experiments are primarily carried out with LK01003 that differs from GSK’872 only in the addition of a cyclopentane ring, in order to explore the significant effect caused by the introduction of this group relative to GSK’872.

a–d Cell viability of L929 cells exposed to various concentrations of GSK’872 (a), LK01002 (b), LK01003 (c), and LK01045 (d), with (open triangle) and without (open circle) the addition of zVAD. e–h Cell viability of L929 cells with RIPK3 knockout (L929RIPK3-/-) exposed to various concentrations of GSK’872 (e), LK01002 (f), LK01003 (g), and LK01045 (h), with (open triangle) and without (open circle) the addition of zVAD. i The activity of caspase 3 in L929 cell lysates treated with 10 µM GSK’872 (magenta solid circle), 5 µM GSK’872 (pink open circle), 50 µM LK01002 (blue open inverted triangle), 50 µM LK01003 (purple open diamond), 50 µM LK01045 (blue open square), or DMSO (black open triangle). j Western blot analysis of RIPK3 co-immunoprecipitation with RIPK1 in L929 cells treated with GSK’872, LK01002, LK01003, or LK01045. Cells are treated with inhibitors at 10 µM for 5 hours. Normalized quantification plots represent the mean ± SD of three independent experiments. k Western blot analysis of caspase 3 and caspase 8 cleavage in L929 cells treated with GSK’872, LK01002, LK01003, or LK01045 for 3 hours. Data in panels (a–i) are presented as mean ± SD from three independent experiments, each with three technical replicates. Data shown in (j, k) represent one of the three independent experiments, all of which show similar results. Statistical differences in (a–h) are assessed using multiple t-tests, and those in (j) using an unpaired t-test. **q < 0.01, ***q < 0.001, ****q < 0.0001. Source data are provided as a Source Data file.
Selectivity is a key consideration in the development of small-molecule kinase inhibitors. The selectivity profile of LK01003 against a panel of 379 kinases demonstrates significantly enhanced on-target selectivity of LK01003 compared to GSK’872. Specifically, LK01003 exhibits an inhibition rate greater than 65% only against RIPK2 and breast tumor kinase (BRK) at a concentration of 1 μM, yielding an S (35) score of approximately 0.005 (Fig. 4a), whereas at the same concentration, GSK’872 inhibits 13 kinases from the overlapping panel of kinases30 (Supplementary Fig. 2). This highlights the superior selectivity of LK01003 for RIPK3. Additionally, using the ADP-Glo assay, the IC50 values of LK01003 against RIPK2 and BRK are determined to be 48 and 248 nM, leading to the selectivity index (SI) of 24 and 124, respectively. We further test LK01003 at a concentration of 10 μM against the panel of 379 kinases (Supplementary Fig. 3). The results show that LK01003 exhibits over 65% inhibition rate towards 12 kinases, yielding an S(35) score of approximately 0.03. Together, these data demonstrate that LK01003 is a highly specific inhibitor of RIPK3 with potent activity.

a Selectivity profile of LK01003 against 379 kinases assayed at 1 μM. The color gradient represents the inhibitory ratio of LK01003 against kinases: magenta represents 100% inhibition, white represents 50% inhibition, green represents no inhibition (0%), and the gradient colors in between represent varying levels of inhibition. The experiment is carried out in duplicate. b Comparison of interaction patterns of GSK’872 and LK01003 with mRIPK3KD by superimposition of their co-crystal structures. c Enlarged view of the DFG motif and αC-helix in the co-crystal structures of mRIPK3KD bound with GSK’872 and LK01003. GSK’872 and LK01003 are shown as magenta and green sticks. d Specific interactions formed by the cyclopentyl group of LK01003 with mRIPK3KD revealed by the co-crystal structure. e Molecular surface representation of the binding pockets for GSK’872 and LK01003 in the co-crystal structures. The overlapped binding site of the two inhibitors is colored in yellow. The unique binding site for the cyclopentyl group of LK01003 is highlighted in red. f The superimposed conformations of the αC-helix and R69 in mRIPK3KD bound with GSK’872 and LK01003. g Interaction patterns of R69 in the structure of mRIPK3KD in complex with LK01003. Source data are provided as a Source Data file.
Mechanism of action of LK01003 with mRIPK3KD and mRIPK3KD R69H
The co-crystal structure of mRIPK3KD in complex with LK01003 provides valuable insights into the molecular mechanism by which LK01003 mimics the R69H mutation and achieves remarkably high selectivity (Supplementary Table 1). In the complex structure, LK01003 mainly occupies the ATP-binding pocket located between the N-lobe and C-lobe of RIPK3KD (Fig. 4b). Superimposition of the co-crystal structures of mRIPK3KD bound with LK01003 and GSK’872 reveals almost identical binding modes in the binding pocket where LK01003 overlaps well with GSK’872 (Fig. 4b, c). However, the introduction of an additional cyclopentyl group in LK01003 induces a significant conformational change in the αC-helix as well as the DFG motif, pushing them away from the ATP-binding site (Fig. 4b). This conformational shift leads to the disorder of the αC-helix and the formation of a hydrophobic binding site between the αC-helix and the DFG motif. The cyclopentyl group establishes hydrophobic interactions with residues M65, L74, and F162 at this new site (Fig. 4b-d), which not only strengthens the binding of LK01003 with the pocket but also renders the kinase domain retain inactive conformations associated with the disordered αC-helix. This in turn changes the conformation of R69, thereby preventing it from forming stable hydrogen bonds with the main chains of V66 and L76, as seen in the complex structure of mRIPK3KD bound with GSK’872 (Figs. 1c, 4f, g). Therefore, like the R69H mutation, LK01003 effectively impedes the dimerization of mRIPK3KD by destabilizing the ordered structure of the αC-helix and reducing the key hydrogen bonds necessary for the stable dimer of the kinase domain. The mechanism of action of LK01003 with RIPK3KD reinforces the notion that the significant conformational changes of the αC-helix and the DFG motif at the dimer interface induced by residue mutation or ligand binding is able to prevent on-target apoptosis. Moreover, based on the binding mode of LK01003 with mRIPK3KD revealed by the co-crystal structure, it can be categorized as a relatively rare Type I1/2 inhibitor.
Although LK01003 disrupts mRIPK3KD dimerization in solution, the complex of mRIPK3KD with LK01003 seems dimeric in the co-crystal structure. It is similar to the binding of GSK’872 to mRIPK3KD R69H, which also appears predominantly as a monomer in solution but as a dimer in the co-crystal structure. This suggests that the assembly of the kinase domain in solution may differ from those in the crystal, with the latter possibly being influenced by crystallization conditions or crystal packing that helps stabilize the dimeric form. To this end, we determine the co-crystal structure of mRIPK3KD R69H in complex with LK01003 (Supplementary Table 1). As in solution, the complex of mRIPK3KD R69H bound with LK01003 also adopts a monomer in the co-crystal structure. Moreover, the binding mode of LK01003 with mRIPK3KD R69H, as shown in the co-crystal structure, closely resembles its binding mode with mRIPK3KD (Supplementary Fig. 4a). Both co-crystal structures of LK01003 in complex with mRIPK3KD and mRIPK3KD R69H show similar conformational changes in the αC-helix and DFG motif of the kinase domain, when compared to the co-crystal structure of mRIPK3KD in complex with GSK’872 (Fig. 3c and Supplementary Fig. 4b). These findings indicate the dual effects of the R69H mutation and the binding of LK01003 render mRIPK3KD R69H sufficiently stable in a monomeric state, not only in solution but also in crystals.
Link between inhibitor-induced conformational changes and on-target toxicity, and unique features of LK series inhibitors
The comparative structural study of mRIPK3KD in complex with GSK’872 and LK01003 reveals distinct conformational changes in the ATP-binding pocket, suggesting that inhibitor-induced conformational changes may be linked to the ability to induce on-target apoptosis. This observation prompts us to comprehensively investigate the relationship between the conformational changes upon binding of RIPK3 inhibitors and their ability to induce apoptosis. We compile a panel of existing RIPK3 inhibitors, including GSK’87230, GSK’84330, Dabrafenib36, GW’39B37, PP238, TAK-63231, and compound 1839, for the measurement of inhibitory activity against RIPK3 as well as on-target apoptosis in cells and the investigation of the ligand binding modes (Fig. 5a). These compounds inhibit RIPK3 ATPase activity with IC50 values ranging from 2 to 56 nM (Fig. 5b) in the ADP-Glo assay and block TNF-α-induced necroptosis in L929 cells with EC50 values ranging from 1.1 to 31 μM (Fig. 5c). We then evaluate these compounds for on-target apoptosis in L929 cells with or without zVAD. The results show that five of these seven compounds, namely GSK’872, GSK’843, Dabrafenib, GW’39B, and PP2, induce on-target apoptosis at concentrations of 3.1, 12.5, 25, 3.1, and 12.5 µM, respectively, which are about 0.4-4 times their respective EC50 values (Fig. 5a, d and Supplementary Fig. 5). This on-target apoptosis is reversed by the addition of zVAD (Fig. 5d and Supplementary Fig. 5), suggesting that these five RIPK3 inhibitors trigger apoptosis alongside their inhibition of necroptosis. Western blot analysis also confirms the activation of the apoptotic pathway, showing the cleavage of caspases 8 and 3 after treatment with these compounds (Fig. 5e). Conversely, TAK-632 and compound 18 do not exhibit on-target apoptosis or induce caspase cleavage, even at high concentrations of 100 and 50 μM, respectively (Fig. 5d, e and Supplementary Fig. 5), which are 26 and 45 times their respective EC50 values (Fig. 5a, c). Therefore, these two compounds are similar to LK01003 in terms of cellular anti-necroptotic activity but do not induce on-target apoptosis, although their IC50 values are weaker than that of LK01003.

a Chemical structures and inhibitory activities of compounds against RIPK3 ATPase activity (IC50) and necroptosis of L929 cells (EC50L929). The crystal structure of mRIPK3KD in complex with Dabrafenib is not resolved, and therefore, the specific type of inhibition by Dabrafenib is not defined. b, c Representative inhibitory profiles of GSK’872 (red solid circle), GSK’843 (blue solid triangle), Dabrafenib (purple solid diamond), GW’39B (orange open circle), PP2 (olive open square), TAK-632 (brown open triangle), and compound 18 (pink open inverted triangle) against RIPK3, measured by the ADP-Glo assay (b) and the necroptosis assay in L929 cell line (c). Necroptosis in L929 cells is induced by treatment with mouse TNF-α (20 ng/mL) and zVAD (10 μM). d Cell viability of L929 cells exposed to RIPK3 inhibitors in the presence (open triangle) and absence (open circle) of zVAD. The concentrations of the inhibitors used are 3.1 µM for GSK’872, 12.5 µM for GSK’843, 25 µM for Dabrafenib, 3.1 µM for GW’39B, 12.5 µM for PP2, 100 µM for TAK-632, and 50 µM for compound 18. e Western blot analysis of caspases 3 and 8 cleavage by RIPK3 inhibitors in L929 cells. The ADP-Glo assay in panel b is performed in triplicate. Panels c and d show data from three independent experiments, each performed with three technical replicates. Panel e presents representative results from one of three independent experiments, all of which yield similar outcomes. Statistical differences are determined by multiple t-tests. **q < 0.01, ***q < 0.001, q < 0.0001. Source data are provided as a Source Data file.
Next, we determine co-crystal structures of mRIPK3KD in complex with GSK’843, GW’39B, PP2, TAK-632, and compound 18 in addition to GSK’872 (Supplementary Table 1). Superimposition of the six complex structures reveals that all compounds bind into the ATP-binding pocket between the N-lobe and C-lobe (Fig. 6), but only the binding of LK01003 disturbs the ordered structure of the αC-helix and extends into a previously unoccupied hydrophobic region (Figs. 4e, 6a, b). However, besides LK01003, the treatment with TAK-632 and compound 18 also does not induce on-target apoptosis of cells, prompting us to dig deeper into the conformational changes of the kinase domain associated with on-target apoptosis. GSK’872, GSK’843, GW’39B, and PP2, which induce on-target apoptosis, exclusively occupy the front region of the ATP-binding site with similar ligand binding modes, leading mRIPK3KD to adopt an active conformation with the DFG motif locked in an “in” state by a salt bridge formed between residues K51 and D161 (Fig. 6a). Given the active conformation of mRIPK3KD upon inhibitor binding, these four compounds are classified as Type I inhibitors. In contrast, the binding of TAK-632 and compound 18 extends into the backside of the ATP-binding pocket, flipping the DFG motif outward and maintaining mRIPK3KD in an inactive conformation (Fig. 6b). These two compounds are thus categorized as Type II inhibitors. Notably, the binding of LK01003, a Type I1/2 inhibitor, also locks mRIPK3KD in an inactive conformation (Fig. 4b-e). Accordingly, the inactive conformations of mRIPK3KD upon binding of inhibitors appear to be key to eliminating on-target apoptosis in cells. Otherwise, apoptosis occurs alongside the anti-necroptotic activity. This finding provides valuable insights into the structural basis of on-target toxicity associated with certain RIPK3 inhibitors.

a Binding modes of Type I inhibitors (GSK’872, GSK’843, GW’39B, and PP2) to mRIPK3KD. b Binding modes of Type II inhibitors (TAK-632 and compound 18) to mRIPK3KD. c Binding modes of Type I1/2 inhibitors (LK01003 and LK01004) to mRIPK3KD. d Binding mode of a Type I1/2 inhibitor (LK01003) to hRIPK3KD. The N-lobe and C-lobe of the RIPK3KD are depicted as yellow and light green cartoons, respectively. The activation loop (AC loop) and αC-helix are displayed as light navy, and orange cartoons, respectively. The residues K51, E61, D161, and F162 in mRIPK3KD and K50, E60, D160, and F161 in hRIPK3KD are shown as sticks. GSK’872, GSK’843, GW’39B, PP2, TAK-632, compound 18, LK01003, and LK01004 are represented by magenta, olive, orange, teal, white, slate, green, and brown sticks, respectively. Hydrogen bonds are marked by dashed lines.
As Type I1/2 inhibitors like LK01003 and LK01045 can disrupt the dimerization of mRIPK3KD, we performed gel-filtration analysis to investigate whether Type II inhibitors have the same effect. The results show that, upon the addition of compound 18, mRIPK3KD is eluted at a position corresponding to the molecular weight of a dimer, similar to the DMSO control, indicating no effect of dimerization disruption found with compound 18 (Supplementary Fig. 6). Similarly, mRIPK3KD is also eluted as a dimer upon the addition of TAK-632 (Supplementary Fig. 6). These results indicate that disruption of dimerization caused by the inhibitors is not a prerequisite for their ability to prevent on-target apoptosis.
In addition, to investigate whether the unique binding pose of LK01003 with mRIPK3KD is also applicable to hRIPK3KD, we solve the co-crystal structure of hRIPK3KD in complex with LK01003 (Supplementary Table 1). The binding modes of LK01003 in two co-crystal structures of mRIPK3KD and hRIPK3KD are almost identical (Fig. 6c, d). The cyclopentyl group of LK01003 also causes a significant shift in the αC-helix and the DFG motif, pushing them away from the ATP-binding site (Fig. 6a, d and Supplementary Fig. 7a) and stabilizing an inactive conformation of hRIPK3KD. This shift also triggers substantial conformational changes in the activation loop (Fig. 6d) and exposes a previously unrecognized hydrophobic binding site between the αC-helix and the DFG motif. The cyclopentyl group establishes hydrophobic interactions with residues M64, L73, and F161 at this new site (Supplementary Fig. 7b), similar to the interactions formed with mRIPK3KD (Fig. 4d). Therefore, the co-crystal structures show that the unique binding mode of LK01003 in mRIPK3KD is also present in hRIPK3KD.
A potential strategy to eliminate on-target apoptosis
To verify the effectiveness of targeting the newly discovered hydrophobic site to avoid on-target toxicity, we conduct the structure-based design of new scaffold RIPK3 inhibitors based on PP2 and Zharp-99 which show significant on-target apoptosis in cells. By superimposing the co-crystal structures of RIPK3KD in complex with multiple inhibitors, we observe that the phenyl ring of PP2 aligns well with the phenyl ring of GSK’872 (Supplementary Fig. 8a). On this basis, we design and synthesize two additional compounds, LK01004 and LK01005, in order to occupy the newly identified site as LK01003 does (Supplementary Fig. 8b). These new compounds demonstrate potent inhibition against RIPK3 in the ADP-Glo assay, with IC50 values of 41 and 81 nM, respectively. Moreover, they exhibit EC50 values of 3.0 and 2.1 μM, respectively, in L929 cells, where necroptosis is induced by treatment with mouse TNF-α and zVAD, confirming their cellular potency against necroptosis by inhibiting RIPK3. To investigate whether these compounds induce on-target toxicity, we perform the cell viability assay. Remarkably, unlike PP2, the CC50 values of LK01004 and LK01005 do not change significantly in the presence or absence of zVAD, indicating that these new compounds do not induce on-target apoptosis in cells (Supplementary Fig. 8c, d). Furthermore, Western blot analysis provides the compelling evidence that at a concentration of 100 μM, LK01004 and LK01005 do not activate the apoptotic pathway, whereas PP2 causes the cleavage of caspases 8 and 3 at a concentration of 25 μM (Supplementary Fig. 8e).
The co-crystal structure of mRIPK3KD in complex with LK01004 is successfully determined at a resolution of 2.78 Å (Supplementary Table 1). A comparable structural analysis shows that the regions shared between LK01004 and PP2 adopt a similar binding conformation (Fig. 6a, c). The newly introduced isobutyl group in LK01004 with flexibility occupies the same binding site as the cyclopentyl group in LK01003, resulting in the outward displacement of the αC-helix and significant conformational changes in the DFG motif (Fig. 6c). In this co-crystal structure, we successfully resolve the structure of the activation loop following the DFG motif. When compared to the co-crystal structure of mRIPK3KD bound with PP2, the activation loop also undergoes substantial conformational changes (Fig. 6a, c).
The selectivity profile of LK01004 at 10 µM is assessed against a panel of 379 kinases (Supplementary Fig. 3). The results demonstrate that LK01004 displays an S (35) selectivity score of approximately 0.18, which is significantly lower than the reported S(35) score of 0.41 for PP240, indicating a significant improvement in kinase selectivity for LK01004 compared to PP2. It should be noted that both LK01005 and LK01004 still exhibit a substantial portion of RIPK3-independent cell killing (Supplementary Fig. 8a, d), likely due to relatively inferior kinase selectivity compared to LK01003, which leads to off-target toxicity.
Analogously, we introduced a cyclopentyl group into Zharp-99 to occupy this miraculous binding site, yielding the new compound LK01033 (Supplementary Fig. 9a). Zharp-9941, a compound derived from GSK’872, is reported to exhibit enhanced anti-necroptotic activity compared to GSK’872, but it triggered stronger on-target apoptosis. As expected, LK01033 shows strong activity in inhibiting necroptosis in both L929 and HT29 cells induced by TNF-α and zVAD, with EC50 values of 0.39 µM and 0.11 µM, respectively, and for reference, the EC50 values of Zharp-99 are 0.07 µM and 0.22 µM, respectively (Supplementary Fig. 9a). Importantly, LK01033 does not induce on-target apoptosis, even at a concentration of 10 µM, whereas 0.1 µM Zharp-99 activates the apoptotic pathway as evidenced by cleavage of caspase 8 and caspase 3 (Supplementary Fig. 9b). The CC50 value of LK01033 in L929 cells does not change significantly in the presence or absence of zVAD, while for Zharp-99 it differs by ~36-fold (Supplementary Fig. 9c, d).
The success of these two additional cases highlights the robustness of the strategy that occupies the newly identified site to eliminate on-target apoptosis while reducing off-target effects on other kinases, paving the way for the development of highly safe and selective RIPK3 inhibitors.
link






