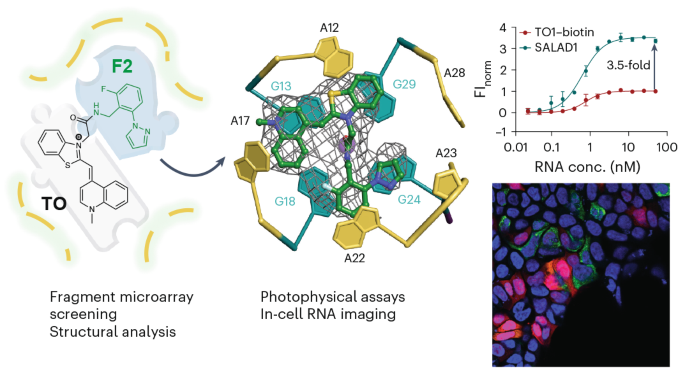
Structure-informed design of an ultrabright RNA-activated fluorophore
Yin, P., Kuang, S. & Nie, Z. Fluorescent RNA tags for in situ RNA imaging in living cells. Anal. Sens. 3, e202200090 (2023).
Google Scholar
Trachman, R. J. & Ferre-D’Amare, A. R. Tracking RNA with light: selection, structure and design of fluorescence turn-on RNA aptamers. Q. Rev. Biophys. 52, e8 (2019).
Google Scholar
Renaud de la Faverie, A., Guedin, A., Bedrat, A., Yatsunyk, L. A. & Mergny, J. L. Thioflavin T as a fluorescence light-up probe for G4 formation. Nucleic Acids Res. 42, e65 (2014).
Google Scholar
Armitage, B. A. Imaging of RNA in live cells. Curr. Opin. Chem. Biol. 15, 806–812 (2011).
Google Scholar
Neubacher, S. & Hennig, S. RNA structure and cellular applications of fluorescent light-up aptamers. Angew. Chem. Int. Ed. 58, 1266–1279 (2019).
Google Scholar
Swetha, P., Fan, Z., Wang, F. & Jiang, J. H. Genetically encoded light-up RNA aptamers and their applications for imaging and biosensing. J. Mater. Chem. B 8, 3382–3392 (2020).
Google Scholar
Huang, K. et al. Structure-based investigation of fluorogenic Pepper aptamer. Nat. Chem. Biol. 17, 1289–1295 (2021).
Google Scholar
Passalacqua, L. F. M., Banco, M. T., Moon, J. D., Li, X. & Jaffrey, S. R. Ferre-D’Amare AR. Intricate 3D architecture of a DNA mimic of GFP. Nature 618, 1078–1084 (2023).
Google Scholar
Paige, J. S., Wu, K. Y. & Jaffrey, S. R. RNA mimics of green fluorescent protein. Science 333, 642–646 (2011).
Google Scholar
Tan, X. et al. Fluoromodules consisting of a promiscuous RNA aptamer and red or blue fluorogenic cyanine dyes: selection, characterization and bioimaging. J. Am. Chem. Soc. 139, 9001–9009 (2017).
Google Scholar
Ji, R. et al. RNA condensate as a versatile platform for improving fluorogenic RNA aptamer properties and cell imaging. J. Am. Chem. Soc. 146, 4402–4411 (2024).
Google Scholar
Dou, C. X. et al. Genetically encoded dual-color light-up RNA sensor enabled ratiometric imaging of microRNA. Anal. Chem. 93, 2534–2540 (2021).
Google Scholar
Bühler, B. et al. Avidity-based bright and photostable light-up aptamers for single-molecule mRNA imaging. Nat. Chem. Biol. 19, 478–487 (2023).
Google Scholar
Chen, W., Zhao, X. Y., Yang, N. Y. & Li, X. Single mRNA imaging with fluorogenic RNA aptamers and small-molecule fluorophores. Angew. Chem. Int. Ed. 62, e202209813 (2023).
Google Scholar
Robinson, J. et al. Cellular visualization of G-quadruplex RNA via fluorescence- lifetime imaging microscopy. J. Am. Chem. Soc. 146, 1009–1018 (2024).
Google Scholar
Su, Y. & Hammond, M. C. RNA-based fluorescent biosensors for live cell imaging of small molecules and RNAs. Curr. Opin. Biotechnol. 63, 157–166 (2020).
Google Scholar
Manna, S. et al. Systematic mutation and unnatural base pair incorporation improves riboswitch-based biosensor response time. ACS Sens. 8, 4468–4472 (2023).
Google Scholar
Dolgosheina, E. V. et al. RNA Mango aptamer-fluorophore: a bright, high-affinity complex for RNA labeling and tracking. ACS Chem. Biol. 9, 2412–2420 (2014).
Google Scholar
Autour, A. et al. Fluorogenic RNA Mango aptamers for imaging small non-coding RNAs in mammalian cells. Nat. Commun. 9, 656 (2018).
Google Scholar
Trachman, R. J. 3rd et al. Crystal structures of the Mango-II RNA aptamer reveal heterogeneous fluorophore binding and guide engineering of variants with improved selectivity and brightness. Biochemistry 57, 3544–3548 (2018).
Google Scholar
Trachman, R. J. et al. Structure and functional reselection of the Mango-III fluorogenic RNA aptamer. Nat. Chem. Biol. 15, 472–479 (2019).
Google Scholar
Trachman, R. J. III et al. Structure-guided engineering of the homodimeric Mango-IV fluorescence turn-on aptamer yields an RNA FRET pair. Structure 28, 776–785 e773 (2020).
Google Scholar
Gotrik, M. et al. Direct selection of fluorescence-enhancing RNA aptamers. J. Am. Chem. Soc. 140, 3583–3591 (2018).
Google Scholar
Ryckelynck, M. Development and applications of fluorogen/light-up RNA aptamer pairs for RNA detection and more. Methods Mol. Biol. 2166, 73–102 (2020).
Google Scholar
Cawte, A. D., Unrau, P. J. & Rueda, D. S. Live cell imaging of single RNA molecules with fluorogenic Mango II arrays. Nat. Commun. 11, 1283 (2020).
Google Scholar
Bychenko, O. S. et al. Red light-emitting short Mango-based system enables tracking a mycobacterial small noncoding RNA in infected macrophages. Nucleic Acids Res. 51, 2586–2601 (2023).
Google Scholar
Lu, X. et al. Symmetry breaking of fluorophore binding to a G-quadruplex generates an RNA aptamer with picomolar KD. Nucleic Acids Res. 52, 8039–8051 (2024).
Google Scholar
Falese, J. P., Donlic, A. & Hargrove, A. E. Targeting RNA with small molecules: from fundamental principles towards the clinic. Chem. Soc. Rev. 50, 2224–2243 (2021).
Google Scholar
Childs-Disney, J. L. et al. Targeting RNA structures with small molecules. Nat. Rev. Drug Discov. 21, 736–762 (2022).
Google Scholar
Yazdani, K. et al. Machine learning informs RNA-binding chemical space. Angew. Chem. Int. Ed. 62, e202211358 (2023).
Google Scholar
Warner, K. D., Hajdin, C. E. & Weeks, K. M. Principles for targeting RNA with drug-like small molecules. Nat. Rev. Drug Discov. 17, 547–558 (2018).
Google Scholar
Bancet, A. et al. Fragment linking strategies for structure-based drug design. J. Med. Chem. 63, 11420–11435 (2020).
Google Scholar
Li, Q. Application of fragment-based drug discovery to versatile targets. Front. Mol. Biosci. 7, 180 (2020).
Google Scholar
Koehn, J. T., Felder, S. & Weeks, K. M. Innovations in targeting RNA by fragment-based ligand discovery. Curr. Opin. Struct. Biol. 79, 102550 (2023).
Google Scholar
Binas, O. et al. 19F NMR-based fragment screening for 14 different biologically active RNAs and 10 DNA and protein counter-screens. ChemBioChem 22, 423–433 (2021).
Google Scholar
Sreeramulu, S. et al. Exploring the druggability of conserved RNA regulatory elements in the SARS-CoV-2 genome. Angew. Chem. Int. Ed. 60, 19191–19200 (2021).
Google Scholar
Cressina, E., Chen, L., Abell, C., Leeper, F. J. & Smith, A. G. Fragment screening against the thiamine pyrophosphate riboswitchthiM. Chem. Sci. 2, 157–165 (2011).
Google Scholar
Zeller, M. J. et al. SHAPE-enabled fragment-based ligand discovery for RNA. Proc. Natl. Acad. Sci. USA 119, e2122660119 (2022).
Google Scholar
Arney, W. & Weeks, K. M. RNA-ligand interactions quantified by surface plasmon resonance with reference subtraction. Biochemistry 61, 1625–1632 (2022).
Google Scholar
Tam, B. et al. Discovery of small-molecule inhibitors targeting the ribosomal peptidyl transferase center (PTC) of M. tuberculosis. Chem. Sci. 10, 8764–8767 (2019).
Google Scholar
Meyer, S. M. et al. DNA-encoded library screening to inform design of a Ribonuclease Targeting Chimera (RiboTAC). J. Am. Chem. Soc. 144, 21096–21102 (2022).
Google Scholar
Connelly, C. M., Abulwerdi, F. A. & Schneekloth, J. S. Jr. Discovery of RNA binding small molecules using small molecule microarrays. Methods Mol. Biol. 1518, 157–175 (2017).
Google Scholar
Wurth, C., Grabolle, M., Pauli, J., Spieles, M. & Resch-Genger, U. Relative and absolute determination of fluorescence quantum yields of transparent samples. Nat. Protoc. 8, 1535–1550 (2013).
Google Scholar
Zhu, Y. et al. Structural modification of nonspecific thiazole orange for ligand-DNA interaction study: understanding the ligand recognition selectivity towards G4-DNA over duplex-DNA. J. Luminesc. 226, 117488 (2020).
Google Scholar
Leonarski, F., D’Ascenzo, L. & Auffinger, P. Nucleobase carbonyl groups are poor Mg2+ inner-sphere binders but excellent monovalent ion binders—a critical PDB survey. RNA 25, 173–192 (2019).
Google Scholar
Fan, Z., Dou, C. X., Tang, L. J., Wang, F. & Jiang, J. H. Genetically encoded RNA sensors for ratiometric and multiplexed imaging of small molecules in living cells. Anal. Chem. 95, 14455–14464 (2023).
Google Scholar
Lu, X., Kong, K. Y. S. & Unrau, P. J. Harmonizing the growing fluorogenic RNA aptamer toolbox for RNA detection and imaging. Chem. Soc. Rev. 52, 4071–4098 (2023).
Google Scholar
Chen, X. et al. Visualizing RNA dynamics in live cells with bright and stable fluorescent RNAs. Nat. Biotechnol. 37, 1287–1293 (2019).
Google Scholar
Yang, M. et al. Targeting a noncanonical, hairpin-containing G-quadruplex structure from the MYCN gene. Nucleic Acids Res. 49, 7856–7869 (2021).
Google Scholar
Calabrese, D. R., Connelly, C. M. & Schneekloth, J. S. Jr. Ligand-observed NMR techniques to probe RNA-small molecule interactions. Methods Enzymol. 623, 131–149 (2019).
Google Scholar
Trachman, R. J. III, Link, K. A., Knutson, J. R. & Ferre-D’Amare, A. R. Characterizing fluorescence properties of turn-on RNA aptamers. Methods Mol. Biol. 2568, 25–36 (2023).
Google Scholar
Winter, G. xia2: an expert system for macromolecular crystallography data reduction. J. Appl. Crystallogr. 43, 186–190 (2010).
Google Scholar
Winter, G. et al. DIALS: implementation and evaluation of a new integration package. Acta Crystallogr. D Struct. Biol. 74, 85–97 (2018).
Google Scholar
Kabsch, W. Xds. Acta Crystallogr. D Biol. Crystallogr. 66, 125–132 (2010).
Google Scholar
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Google Scholar
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Google Scholar
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Google Scholar
Krissinel, E. & Henrick, K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 372, 774–797 (2007).
Google Scholar
link






